|
6/24/2023 0 Comments Samtools threads 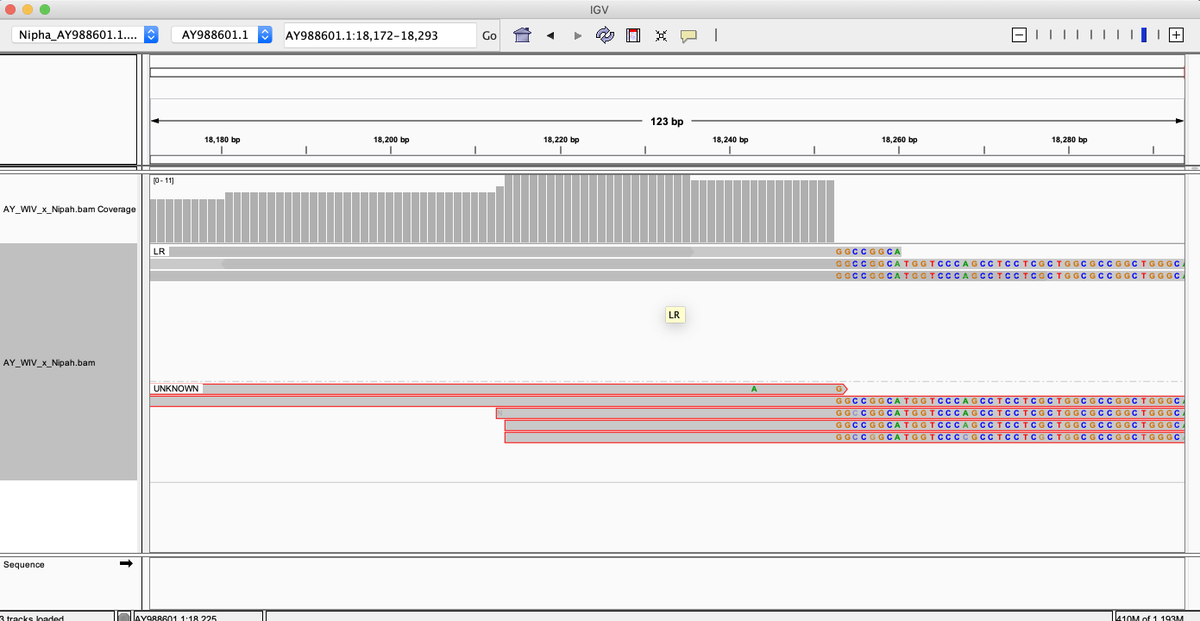
This is indicated under every history item with e.g.: "database: hg19" for a link to hg19, or "database: ?" if the link is missing. The alignment files have to be linked to a reference genome by galaxy. 
Satmools accepts sequencing alignments in the same, either SAM or BAM format ( ). If fewer threads are desired, they can be specified with the -jobs flag, e.g. However, since the project seems to lack support and contains fatal bugs this project was forked at: Where mybams.fofn is a file of BAM files, and genome.fa.fai is the output of samtools faidx or alternately a newline separated list of chromosomes. Git clone git:///samtools/samtools.gitīecause samtools does not support parallization of the mpileup command, the project was forked to include paralellization support: You can check out the most recent source code from the github project page with: The source code releases are available from the download page. SAM Tools provide various utilities for manipulating alignments in the SAM format, including sorting, merging, indexing and generating alignments in a per-position format. Is simple enough to be easily generated by alignment programs or converted from existing alignment formats Īllows most of operations on the alignment to work on a stream without loading the whole alignment into memory Īllows the file to be indexed by genomic position to efficiently retrieve all reads aligning to a locus. 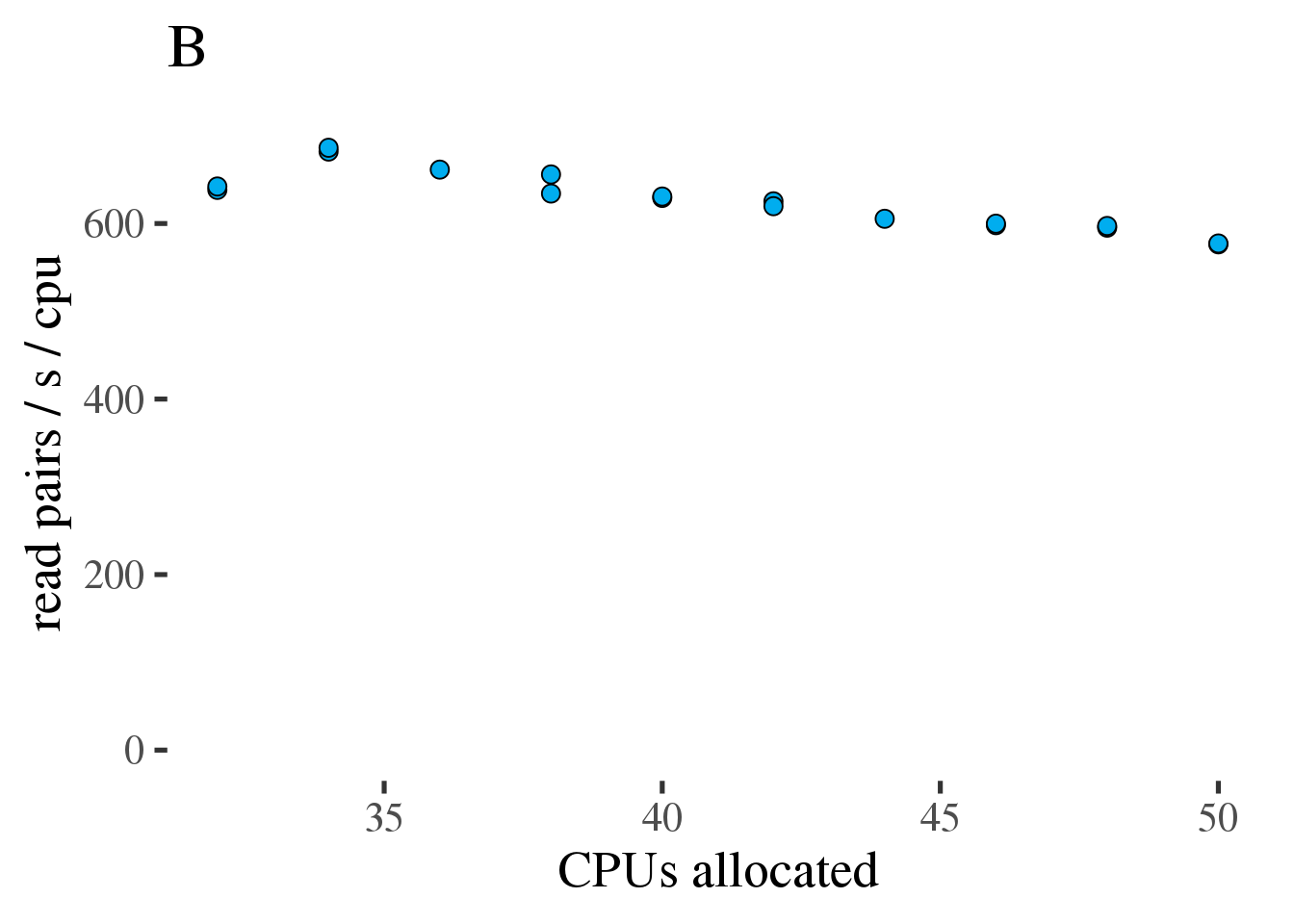
Is flexible enough to store all the alignment information generated by various alignment programs SAM (Sequence Alignment/Map) format is a generic format for storing large nucleotide sequence alignments. Samtools mpileup (supporting parallelization)
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
